Central Control of Feeding Behaviour and Energy Expenditure (C2OFFEE)
Head: Serge Luquet (DR CNRS)
In mammals, the regulation of energy homeostasis is finely regulated. In fact, despite fluctuating, blood glucose levels, body weight, and fat contents remain within narrow ranges and experimentally-induced perturbations invariably result in a rapid return to ‘set points’ when normal conditions are restored. To accomplish this, circulating peripheral factors, such as hormones (e.g., insulin, leptin, ghrelin) and nutrients (e.g., glucose, lipids), promptly modulate discrete neural circuits that trigger changes in the basal metabolic rate and/or feeding behaviours. Disruption of these circuits can give rise to life-threatening conditions that include metabolic diseases such as obesity and diabetes in both humans and rodent models. Therefore, it is crucial to understand the key mechanisms underlying the proper equilibrium between energy intake and expenditure. The core approach of our team Central COntrol oF Feeding behaviour and Energy Expenditure (C2OFFEE) is to leverage the power of modern molecular genetic tools in integrated approaches in order to dissect in vivo the physiological, cellular and molecular basis by which the dialogue between the brain and peripheral organs result in adaptive regulation of the various parameters of energy balance notably in its rewarding, motivational and sensory components, metabolic efficiency, allostatic response.
Both homeostatic and reward circuits intimately collaborate in gating adaptive responses to changes in nutrients availability and are, directly and indirectly, controlled by extrinsic signals (sensory cues, including odors), cognitive factors (stress, environmental cues and stimuli), intrinsic factors including circulating energy-related signals (hormones, nutrient) and afferent nervous inputs from the periphery (gut-brain vagal axis). It is now established that the modern food environment has a drastic impact on the brain’s ability to control proper feeding and energy expenditure. While overconsumption is clearly identified as a primary culprit, a large body of evidence supports that the development of obesity and its related disorders result from an interaction between genes and environment. Our goal is to explore the physiopathological bases of these processes and identify molecular components involved in the establishment of susceptibility/resilience to reward- and metabolic-related disorders. 4 research axes are developed under this common interest (see schematic representation and detailed methodology).
Main axes
Axis 1: Impact of metabolic state on multisensory signaling.
Axis 2: Astrocyte ‘control of metabolism
Axis 3: Molecular determinants of brain susceptibility/resilience to feeding & metabolic diseases.
Axis 4: Interoceptive dynamics controlling homeostatic and reward processes
Legend
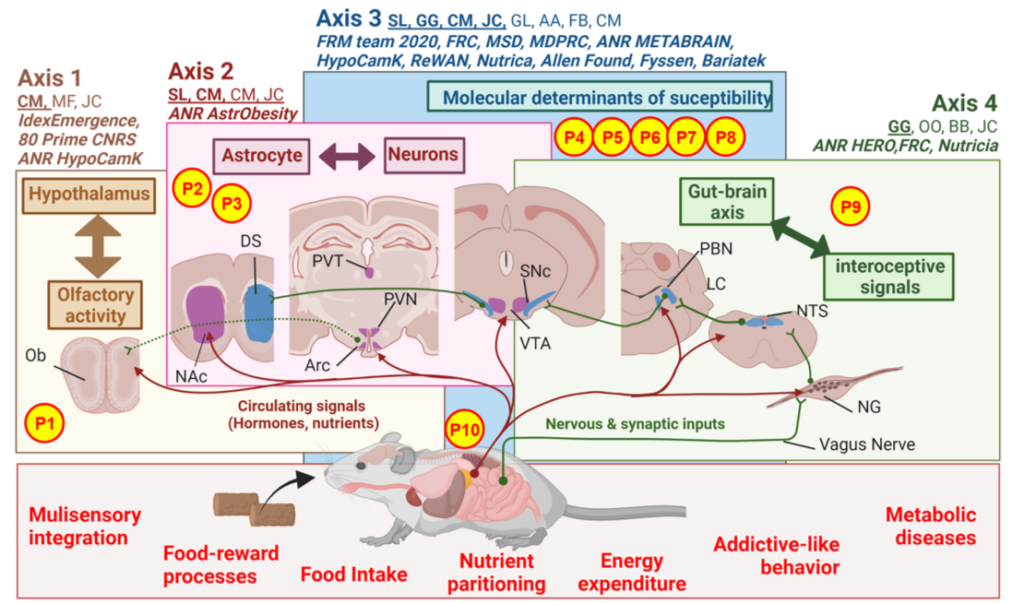
The C2OFFEE team main research axes and work force. Circulating (red) and nervous/synaptic signals (green) conveying energy states are integrated at various levels in the brain and reflect the bimodal dialogue between the brain and peripheral organs. Through 4 interdependent research axes the team explores: (1) the connection between hypothalamic encoding of hunger and multisensory/olfaction capacity, (2) the role of astrocytes in the hypothalamus and reward system in long-term adaptations to an obesogenic environment, (3) the genetic and molecular determinants that siege at brain susceptibility or resilience to the detrimental action of modern-food environment and (4) the functional networks and their cellular/molecular underpinnings trough which interoceptive signals and the extended gut-brain axis control metabolic efficiency, reward processes and addictive-like behaviors.
Work force: Serge Luquet (SL), Giuseppe Gangarossa (GG), Claire Martin (CM), Julien Castel (JC), Guangping Li (GL), Anthony Ansoult (AA), Fanny Bain (FB), Caroline Leger (CL), Maya Faour (MF), Oriane Onimus (OO), Nour Mesto (NMe), Benoit Bertrand (BB), Nejmeh Mashhour (NMa), Camille De Almeida (CdA), Claudia Grajeda (CG)
Abbreviation: Nodose Ganglia (NG), Arcuate nucleus (Arc), Paraventricular thalamus (PVT), Paraventricular nucleus (PVN), Nucleus Tractus Solitarius (NTS), Olfactory bulb (Ob), Ventral Tegmental Area (VTA), Substantia Nigra compacta (SNc), Dorsal Striatum (DS), Nucleus Accumbens (NAc), Parabrachial nucleus (PBN), Locus Coeruleus (LC)